Difference between revisions of "HiSeq2000 - Next Level Hacking"
(→Comands) |
(→Comands) |
||
| Line 589: | Line 589: | ||
| | | | ||
| Move filters / sensors ? | | Move filters / sensors ? | ||
| + | |- | ||
| + | |- | ||
| + | | | ||
| + | | | ||
| + | | | ||
| + | | | ||
|- | |- | ||
Revision as of 16:34, 4 April 2018
We got a HiSeq 2000, Next Level Sequencing Machine from the Genomics Facility of Department of Biosystems Science and Engineering in Basel. Contact through Biozentrum, University of Basel. We got it for free with the only disclaimer: "The biohackers should understand that they are responsible to organize and pay for the transport as well as that there is no warranty or support that can be given neither by us nor the DBSSE."
This type of machine seems to be quite difficult to get up and running and also reagents, flowcell-kits and software licences can be expensive. Since more of these machines seem to show up in second hand (there are new machine generations by Illumina) it would be worth trying to find a way to make them work. Sequencing for all.
Specifications:
https://www.illumina.com/documents/products/datasheets/datasheet_hiseq2000.pdf
The HiSeq2000 (200Gb) was introduced in the year 2010. Followed by HiSeq2500 (500Gb) in 2012. And HiSeq X Ten (1000Gb) in 2014. In 2017 the NovaSeq series of machines was launched.
The machine is a quite early on, from March 2011, Serial Number is 700792, so the machine can not be updated to software and chemistry v4. Only machines with SN#
7001403 or higher can get the FPGA update v4.

Contents
First Inspection
I made a first inspection on the machine. It seems very well made (2011). I still think it would be cool to make it run as is. It's basically a big microfluidic system. So if we get the pumps and the cameras to work we can hack it into anything we want 🙂. Even if it's not for sequencing - it's basically a holder for flow-cells with a fluorescence camera attached to it. And 32 channels with pumps and selector valves that attach to the flow cells. Plus a fridge and a computer.
And peltier for heating and cooling (pcr). Now trying to get the control software. I also think the system is "relatively" open... the software can be downloaded and kind of installs, there is no ID checking on the supplies or anything. Looks very hackable. Also all the cases can be opened easily. Let's do a weekend hack-session on it.
I think it's great opportunity to learn about next level sequencing and about how theses machines work.
Fluidic System:
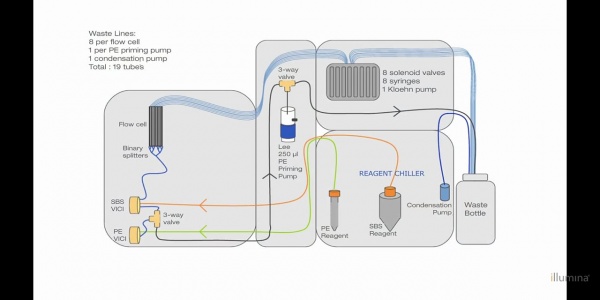
Some pictures from the inside of the machine:
Lausanne Bio-Hackerspace Hackuarium got a HiSeq2000 (SN# 700918) and Gustavo dissected it. Here some pictures that Rachel sent me with comments form what I think components are:
Chemistry
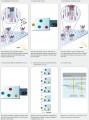
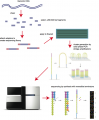
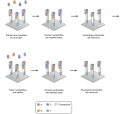
Some images describing the Illumina Next-Generation Sequencing Chemistry:
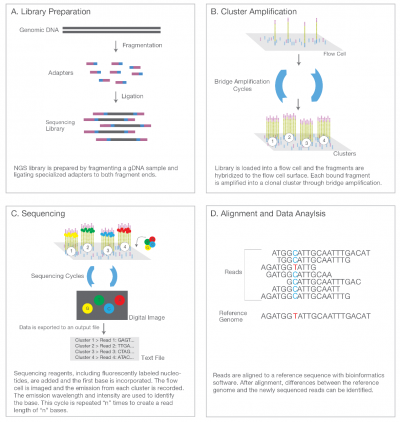
Illumina uses a process called "Sequencing-by-synthesis"
The HiSeq (and MiSeq) use 4-colour SBS
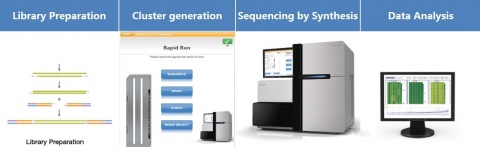
The full DNA to Data solution:

Paper on the 4 color SBS:
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1702316/
Video with the possibilities for library preparation:
https://www.youtube.com/watch?v=_yC0Bzw3WbQ
Library prep kit with sample purification by magnetic beads:
https://www.youtube.com/watch?v=UE1TAZZPUFI
The CBot 2 System is used to prepare the clusters on the flow cell:
https://emea.illumina.com/products/by-type/accessory-products/cbot.html
HiSeq (and TrueSeq) Rapid Cluster Kits can be clustered on the HiSeq (on-board cluster). They also seem to have a lower read and are cheaper:
https://emea.illumina.com/products/by-type/sequencing-kits/cluster-gen-sequencing-reagents/hiseq-rapid-cluster-kit-pe-sr.html
Chemisty Kits Option (running on our machine):
All Library Prep Kits
Cluster Generation (cBot needed):
- TruSeq SR Cluster Kit v3 - cBot-HS - Price: 4360 CHF
- TruSeq PE Cluster Kit v3 - cBot-HS - Price: 6674 CHF
- TruSeq Rapid SR and PE Cluster Kits, cBot Duo Cluster Kit - Price: Request
Maybe not compatible (for reference):
- HiSeq SR Rapid Cluster Kit v2, Price: 1002 CHF
- HiSeq PE Rapid Cluster Kit v2, Price: 1540 CHF
SBS Reagents:
- TruSeq SBS Kit v3 - HS (50 cycles), Price: 2552 CHF
- TruSeq Rapid SBS Kit - HS, Price: Request
Maybe not compatible (for reference):
- HiSeq Rapid SBS Kit v2 (50 cycles), Price: 587 CHF
Question:
- Are the "HiSeq Rapid Cluster Kit v2" compatible with HiSeq (before the v4 update). Or are the "TrueSeq Rapid Cluster Kits"?
- What Kit can be clustered on our HiSeq?
Software
Our machine with SN# 700792 can not be updated to HiSeq Control Software (HCS) v2.2.37 or higher.
Possible options are (depends on software on the FGPA, how to find out?):
- HCS v1.5, RTA v1.13
- HCS v2.0, RTA v1.17
- HCS v2.2.38, RTA v1.18 - Not compatible
- HCS v2.2.58, RTA v1.18 - Not compatible
On the "do not eject" virtual drive from the machine it looks as if the last update was:
override_2013-08-21__05_11_55.cfg
So that was just after releas of HCS 2.0
The HiSeq instrument computer employs 64-bit Windows Vista.
Question:
- Does V2.0 have a Rapid Run mode?
- What is the "Include_Override.cfg" file?
Download for hcs_1-5-15-1:
https://support.illumina.com/downloads/hcs_1-5-15-1_rta_1-13_sav_1-8_software.html
Download for hcs_2_0_12:
https://support.illumina.com/downloads/hcs_2_0_12.html
Workstation (Computer)
Dell Precision T7500 Tower-Workstation
AMD FirePro V3750 256MB
48 Giga Ram

http://euro.dell.com/content/products/productdetails.aspx/workstation-precision-t7500?c=de&l=de&s=corp
http://www.dell.com/support/article/us/en/4/sln291329/precision-t7500-windows-xp-and-windows-vista-driver-install-guide?lang=en#Broadcom57XXGigabitControllerDriver
All the drives were removed from the computer when we got it. There is a raid controller board, an ethernet card and the Phoenix Video Card installed in the machine.

Unfortunately the computer has a problem with the power supply and does not start up properly and we need to press the button on the powersupply to get it going somehow.
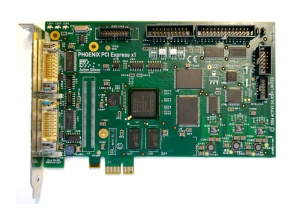 The Phoenix PCI Express x1
The Phoenix PCI Express x1
The screen comes with a touch screeen interface.
Software Installation
We had to completely reinstall the control computer. All the files needed to get the machine up and running could be found (2018) as free downloads on the internet. Here is how:
- First install Windows Vista
- Install the ethernet driver, the grafics card driver and the Chip-Set Driver from the Dell Support Hompage (links below)
- Install the HiSeq Controll software (in our case we guessed Version HCS 2.0, depends on the FPGA version and SN of the machine) using the install.bat
- The connect the HiSeq and turn it on, wait for 2 minutes until all the serial device show up
- Then had to install the FTDI2xx64 driver for the USB serial ports from the FTDI website (for Windows Vista)
- Also had to put a copy of the FTDI2XX.dll in the Illumina/ControlSoftware folder. (why?)
- Install the Camera driver, reboot
- Also put a copy of the camera.dll into the Illumina/ControlSoftware folder (not sure if needed).
- Then the software keeps asking for an "Override" file on C:, so I copied to "Override_xx" file from the "DONOTEJECT" drive of the machine to C: and renamed it
- Also the software needs drives up to E: for data / setting storage (just used some USB sticks)
- Then start the control software and wait for the initialization to complete.
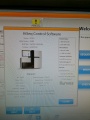
After many tries the successfully initialized machine interface:
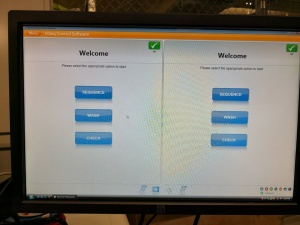
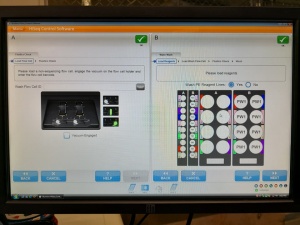
Ready for some sequencing (or further tests)
DCAP-api Driver:
https://dcam-api.com/downloads/
USB / Serial Port Sniffer that can be useful to reverse engineer the communication:
https://freeusbanalyzer.com/
https://docs.microsoft.com/en-us/sysinternals/downloads/portmon
Links and Information
Illumina Next Level Sequencing:
https://www.youtube.com/watch?v=womKfikWlxM&feature=youtu.be
Expert Videos:
https://www.youtube.com/playlist?list=PLKRu7cmBQlai-GUWeAN-eHD5xRcCXDW-D
HiSeq2000 support page:
https://support.illumina.com/sequencing/sequencing_instruments/hiseq_2000.html
HiSeq 2000 User Guide:
http://fantom.gsc.riken.jp/5/sstar/images/1/11/HiSeq2000_UserGuide_15011190_D.pdf
HiSeq Compatibility Chart:
https://support.illumina.com/content/dam/illumina-support/help/version_compatibility/Default.htm
Video of the scanning:
https://www.youtube.com/watch?v=tuD-ST5B3QA
Illumina Support:
Switzerland +41 565800000 +41 800200442
Discussion
The goal is to make it work!
Let's discuss on the forum.. http://forum.hackteria.org/t/hiseq2000-next-level-hacking/325/1
- Where to get/buy the reagents and flow cells
- Hackquarium Lausanne got a similar machine and dissasembled it. We can get pointers from Gustavo on how to take it all apart!
- Muffatto there's a fluorescence microscope inside (afaik), the issue is to reduce it in size and still have it working
- Muffatto: erik from biocurious reverse engineered the chemistry of the system for BGI
- Bengt: Absolutely in on building an openseq kind of thing that can run original reagents. And then re-engineering the system for smaller/cheaper/simpler - even better if combined with an effort to make open reagents - but that is 2 tracks that can progress independently
Technical Descriptions / Findings
The fluorescent readout system with lasers and CCD cameras
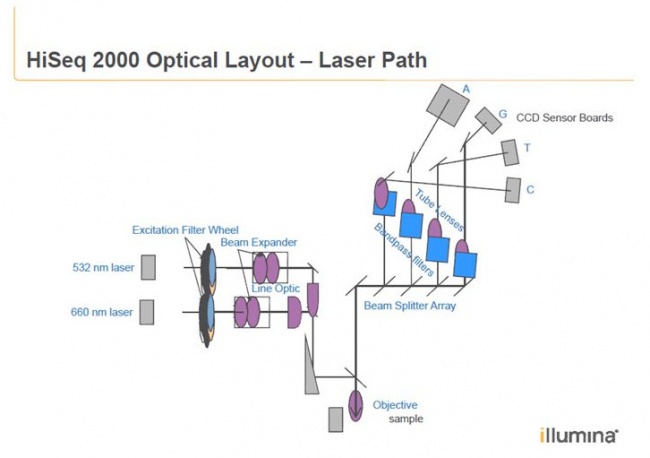
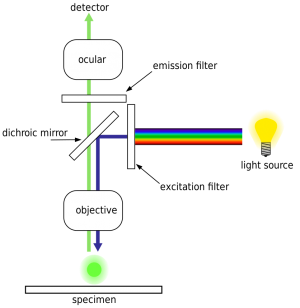
Patent by Illumina:
https://www.google.com/patents/DE202011003570U1?cl=it
The HiSeq uses an epifluorescence microscope design shown in the diagram. Light of the excitation wavelength is focused on the specimen through the objective lens and the fluorescence emitted by the specimen is focused back the detector by the same objective.
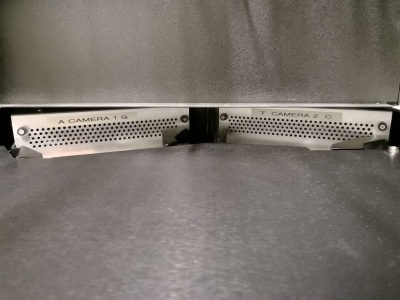
Here you can see the two camera units with even the letters A G T C written on it.

The laser calibration sheets that came with the machine:
The readout system of the HiSeq uses Line Imaging:
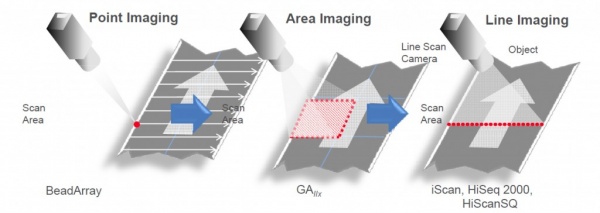
The Flow-Cell
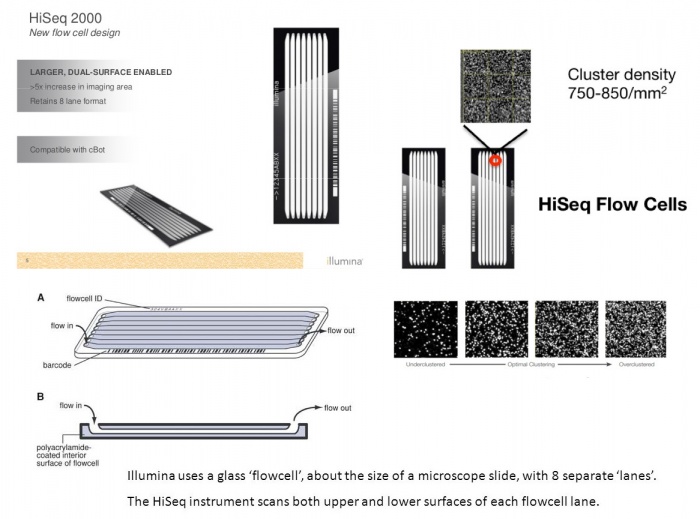

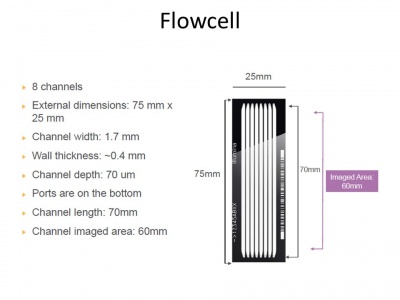
The recommended maximum cluster density is 750,000-850,000 clusters/mm² when using Illumina's v3 cluster generation and sequencing reagents in combination with HCS v1.4. That makes 866²-921² clusters or 1.08-1.12 um² in average per cluster.
Some rough calculations/estimations on what is going on in the machine
The DNA clusters are about 1 Micrometer in size. (or bigger for older machines/software, maybe 2-5 um)
The lines on the flow cells are about 1 mm large what means 1000 clusters. (Or lager, up to 1.7 mm)
The reading speed is about 1mm per second or 1000x1000 clusters per second. Or 1 Mega bases.
The lines are 6 cm long and there are 10 lines per flow cell.And two flow cells.
The means 600 Mega Clusters (Bases) per flowcell per complet run. And that takes about 600 seconds or 10 minutes.
Then flush the flow-cell to add the next base (SBS, sequencing by sythesis). Then start over again.
Unitl the whole 150 bases long DNA sequences are read.
This takes about 4 days... 10 Genoms.
The cluster are red in 4 colors / letters at the same time through 2 lasers exciting 4 colors in fluorescence.
By 4 CCD line cameras with Time delay and integration (TDI). Line cameras with 1 micrometer resolution, 1000 lines a second. Or 1 picture 1000x1000 per second...
The chemistry cost some hundreds to some thousands... but for what it does its not so bad. And you get the chemistry kit with the flowcell. So all the magic and the rest is just some kind of state of the art open hardware 🙂
Hardware components
Nice description on what hardware components are used in the HiSeq on the following page (see comments):
https://blogs.swarthmore.edu/Illumina+GAIIx+Teardown/?p=125
- two lasers (Laserquantum ignis 660nm, gem 532nm with SMD6000 drivers)
- filter revolvers, beam expanders (Linos 2-8x) followed by a barrel lenses for each wavelength, a combiner to join the two excitation wavelengths
- piezo actuator for the Z-Stage (Physik Instrumente P-601 with driver E-601 and E-801 Sensor module)
- Nikon CFI Plan APO VC 20x Objective.
- XY-table (Parker 803-4099, something like the XR400 series, driven by a ViX-250-IH driver module).
- The “Docking Station” with the Flowcells are mounted on three Stepper adjustable points to align them with the focal plane of the line scanner.
- The fluorescence signal is divided by a fixed filter set 4x
- 4 CCD cameras with S10405 line CCDs from Hamamatsu (DIL 40 package).
- Two Hamamatsu Camera Control Boards (Model C10000-509) are each controlling two of these line cameras.
- Illumina board with: line CCDs -> 8 LTC2203 25Msps 16-Bit ADCs -> Altera Cyclone II FPGA. Spartan XC3S4000 FPGA and XC95288 CPLD (both Xilinx)
- Data is collected by the Phoenix AS-PHX-D48CL Frame grabber card in the Computer
Port usage
| Device | Port | Settings |
|---|---|---|
| ARM9BoardSerialPort / ARM9 Chem | Port: COM3; ARM9CHEM ;CM00006 | |
| ARM9BoardDiagSerialPort / ARM9 Diag | Port: COM6; ARM9DIAG | |
| FPGA P1 Command / command_com_port_num | Port: COM10; IL000004 | 115200 baud |
| FPGA P2 Response / response_com_port_num | Port: COM11; IL000005 | 115200 baud |
FlowcellFluidics1 |
||
| FtdiViciValve1 / VICI A1: | Port: COM16; VICIA1 ;CM00004 | 19200 baud |
| FtdiViciValve2 / VICI A2: | Port: COM20; VICIA2 ;CM00043 | 19200 baud |
| KloehnControllerPump / Kloehn A | Port: COM18; KLOEHNA ;CM00001 | |
FlowcellFluidics2 |
||
| FtdiViciValve1 / VICI B1: | Port: COM17; VICIB1 ;CM00002 | 19200 baud |
| FtdiViciValve2 / VICI B2: | Port: COM21; VICIB2 ;CM00044 | 19200 baud |
| KloehnControllerPump / Kloehn B | Port: COM19; KLOEHNB ;CM00003 | |
Laser 1,Green532,Smd6000 |
Port: COM12; IL000006 |
9600 baud |
| Laser 2,Red660,Smd6000 | Port: COM13; IL000007 | 9600 baud |
XMotor_MDrive_5mm (X-Stage) |
Port: COM7; IL000001 |
9600 baud |
| YMotor_VixServoIH_10nm (Y-Stage) / Servo linear motor MT49420 | Port: COM8; IL000002 | 9600 baud |
| Z-Stage | Port: COM9; IL000003 | |
Barcode_Reader |
Port: FTE2V9ML ; COM5 (com_port_num = 4) |
|
| PTC | Port: COM2 (com_port_num = 1) | 9600 baud |
| rs232 | Port: COM10 (offset 9) | |
| PCIO Board | Port: COM4; PCIO (com_port_num) | |
| Test Port | Port: IL000010 | |
| PC Camera Ctrl | Port: COM14; IL000008 | |
Phoenix Cam 1? / AS-PHX-D48CL-PE1 (C10000-509) |
Port: COM22 |
|
| Phoenix Cam 2? / AS-PHX-D48CL-PE1 (C10000-509) | Port: COM23 | |
| Flash Drive | Port: COM15; IL000009 |
Scanner.ChemistryModule
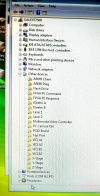
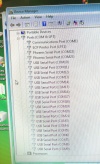
During the software install I got this rather interesting list on the Hardware Manager showing all the device names.
After the installation of the USB serial port driver the names disappeared.
Ports:
- ARM9 Chem
- ARM9 Diag
- Flash Drive
- FPGA P1 Command
- FPGA P2 Response
- Kloehn A
- Kloehn B
- Laser 1
- Laser 2
- Multimedia Video Controller ?
- PC Camera Ctrl ?
- PCIO Board
- Test Port ?
- VICI A1
- VICI A2
- VICI B1
- VICI B2
- X-Stage
- Y-Stage
- Z-Stage
Drives
Data (D:) Data (E:) DVD (F:) Removable (G:) DoNoEject (H:)
Comands
FPGA
| Command | Receive | Parameter | Function |
|---|---|---|---|
| MISCRVC\n | MISCRVC 3.0.14 | Reads version number of a board C | |
| MISCRV\n | MISCRV 3.10.3 | Reads version number of a board | |
| RESET\n | @LOG The FPGA is now online. Enjoy! / RESET | Reset And Cancel All Commmands, is about to reset FPGA | |
| LEDMODEs m\n | LEDMODEs | s=[1,2] m=[0,2,3,4,7] |
Big front LED-bar light mode. s= left and right section (for each flow cell) |
| LEDPULSRATE r\n | LEDPULSRATE | r=[1-255] (milliseconds) | Puls rate of LED-bar. Controls, the frame update rate for the pulsing animation. |
| LEDSWPRATE r\n | LEDSWPRATE | r=[1-255] (milliseconds) | Sweeping rate of LED-bar. Controls, the frame update rate for the sweeping animation. |
| LEDMODETMPST 0\n | LEDMODETMPST | ??? | |
| EM2VL_DN p\n | EM2VL_DN | p=[0..100? / 10,20,40] (percent) | Emission Filter 2, Velocity Percent Down |
| EM2VL_UP p\n | EM2VL_UP | p=[0..100? / 10,20,40] (percent) | Emission Filter 2, Velocity Percent Up |
| EM2RDV_DN\n | EM2RDV_DN v | v = [0..10?] 10 | Emission Filter 2, Read position? |
| EM2RDV_UP\n | EM2RDV_UP v | v = [0..10?] 10 | Emission Filter 2, Read position? |
| EM2RD\n | EM2RD 0 1 0 0 | a b c d =[0,1] / a= , b=inPathSensor , c=OutOfPathSensor , d= / position = IntoPath / OutOfPath / Unknown | |
| EM2O\n EM2I\n EM2RD\n | Move filters / sensors ? | ||
| EXsHM2\n | EXsHM2 | s=[1,2] filter path | Home excitation filter 1 or 2 to laser-safe blocked position. |
| EXsMV p\n | s=[1,2] p=[-71,71] | ?? | |
| EXsVL p\n | s=[1,2] p=[131072] | ?? | |
| EXsCUR p\n | s=[1,2] p=[35] | ?? | |
| TnMOVETO p\n | n = [1,2,3] (motor), p=18504 (steps, 656 steps/mm) / (3025 steps (4.610 mm)? | Tilt motor (n) closed-loop absolute encoder moves. | |
| TnRD\n | TnRD q | n = [1,2,3] (motor) q=steps ?encoder und motor steps equal? | Read tilt motor encoder. |
| DOORSSTATE p\n | DOORSSTATE | p=[0,1] | Set door state ? |
| DOORSOPEN\n | |||
| LASERPB p\n | LASERPB p | p=[0..255/500/or more] (DAC/mW?) | Set laser power. |
| SWLSRSHUT 1\n | SWLSRSHUT | Opening laser shutter | |
| LSEN\n | LSEN | Laser Safety Enable Reporting | |
| ZMV p | ZMV | p=[1311..64224] | Z-Move And Trigger. Move Z-motor with ramp. ZMV will trigger camera immediately and ramp is HW controlled |
| TDIYEWR 6000012\n | TDIYEWR | ? | TDI |
| SWZLAG 25\n | SWZLAG | ||
| EX1HM2\n | LaserShutterArbiter | ||
| EX2HM2\n | |||
| ZADCR\n | ZADCR 1266 | ADC Read | |
| ADC noise = 10, warning limit = 655, error limit = 3277, numSamples = 10x8, avg = 1263, posNoise = 10, negNoise = 10 | |||
| ZDACW 63438\n | ZDACW | ADC Write | |
| ZADCAE 1\n | |||
| ZYT 0 3\n | ZYT | ||
| ZTRG 0\n | ZTRG | trigger | |
| ZSPEC -47 65491\n | |||
| ZSTEP 1288471\n | |||
| ZSEL 0\n | |||
| ZCLK 1\n | |||
| T1CR\n | |||
| T1HM\n | |||
| T1RD\n | T1RD 0 | Tilt Motor 1 | |
| T2CR\n | |||
| T2HM\n | |||
| T2RD\n | T2RD 0 | Tilt Motor 2 | |
| T3CR\n | |||
| T3HM\n | |||
| T3RD\n | T3RD 0 | Tilt Motor 3 | |
| T1CUR 35\n | |||
| T2CUR 35\n | |||
| T3CUR 35\n | |||
| T1VL 62500\n | |||
| T2VL 62500\n | |||
| T3VL 62500\n | |||
| ~DOORSOPEN\n | |||
| ~EM2O\n | |||
| EM2I\n | Emission Filter 12: 'EmissionFilter2' | ||
| EM2O\n | |||
| EM2RD\n | |||
| EM2RDV_DN\n | |||
| EM2RDV_UP\n | |||
| EM2VL_DN 10\n | |||
| EM2VL_DN 40\n | |||
| EM2VL_UP 10\n | |||
| EM2VL_UP 20\n | |||
| Initialize Excitation Filter 2 | |||
| EX1CUR 35\n | |||
| EX1VL 131072\n | |||
| EX2CUR 35\n | |||
| EX2VL 131072\n | |||
| Excitation Filter 1: Velocity = 40 MHz / 131072 = 305.18 Hz | |||
| MIOCOMPR\n | Compensator: Selecting Galvo-type compensator | ||
| MIOGSTEPS1 1 0\n | Excitation Filter 1: Homed to center of sensor in 1292.4745 ms | ||
| MIOGSTEPS30 1 0\n | Compensator: Moved OutOfPath in 420 ms | ||
| MIOGSTEPS30 5000 0\n | |||
| MIOGSTEPS30 5000 2\n | |||
| MIOGVAL 00\n | |||
| MIOGVAL 200 0\n | |||
| MIOGVAL 200 130\n | |||
| MIOMVAL 00\n | |||
| MIOSEL 2\n | |||
| MIOSPIINIT\n | |||
| RESET\n | |||
| SW_BRUNO 1\n | |||
| SWBEADCOMPAT 0\n | |||
| SWCEILING65535\n | |||
| SWCVGAIN1500\n | |||
| SWCVGAIN2500\n | |||
| SWCVHTPX 350\n | |||
| SWCVLIM1 3\n | |||
| SWCVLIM2 7\n | |||
| SWCVOFST 250\n | |||
| SWCVPRT1 0\n | |||
| SWDITH_IGAIN 100\n | |||
| SWDITH_IHIST 4\n | |||
| SWDITH_SHIFT 20\n | |||
| SWDITH_SIZE 100\n | |||
| SWFTLSR 0\n | |||
| SWFTLSR 1\n | |||
| SWLINECOUNT 1\n | |||
| SWLSRSHUT0\n | |||
| SWLSRSHUT1\n | |||
| SWTIME 0\n | |||
| SWTIME 1\n | |||
| SWVIX 0\n | |||
| SWVIX 1\n | |||
| SWYZ_POS 1\n | |||
| SWZLAG 10\n | |||
| SWZLAG 25\n | |||
| TDIYEWR 6000008\n | |||
| TDIBB 0\n | |||
| TDIMASK 24\n | |||
| TDINEW_ATS\n | |||
| TDIZ_PI_STAGE\n | Offaxis Camera Initialize | ||
| TRAHSEN 0\n |
ARM9CHEM
| Command | Parameter | Function |
|---|---|---|
| ?PRETMP\r | 9.007C | |
| ?VSTAT\r | 3:A1\r\n | |
| ?VSENSE:1\r | 4:A1\r\n | |
| ?FCTEMP:1\r | 23.824C:A1\r\n | |
| ?HDDRVER\r | ||
| ?IDN\r | ||
| WASTE:0:1\r | ||
| VACUUM:1\r | A1\r\n | |
| DAC:4:2130\r | ||
| FCRTD:0:M:0.957142\r | ||
| FCTEC:0:0\r | ||
| FCTEMP:0:P:0.2\r | ||
| INIT\r | ||
| RERTD:0:M:1\r | ||
| RETEC:0:P:0.8\r | ||
| RETEMP:2:4.00\r | ||
| STRMASK:2:2\r |
X-Motor
| Command | Receive | Parameter | Function |
|---|---|---|---|
| PR VI | PR VI\r\n410\r\n | 410 (1.0009765625 mm/sec) | Parameter Read velocity |
| PR VM | PR VM\r\n6144\r\n | 6144 (15 mm/sec) | Parameter Read velocity |
| VM=4096 | VM=4096\r\n | ||
| VI=1 | VI=1\r\n | Setting velocity for finding negative limit sensor to 10.00 mm/sec | |
| MR -409190 | MR -409190\r\n | -409190 to negative limit | moving and read ? |
| Xmit | Recv | ||
| DE=1\n | DE=1\r\n> | ||
| EX 1\n | exit? | ||
| MA -4915\n | MA -4915\r\n> | ||
| MR -409190\r | |||
| PM=0\n | PM=0\r\n> | Homing part 3 finished OK | |
| PR ER\n | PR ER\r\n0\r\n> | ||
| PR ER\r | PR ER\r\n84\r\n> | ||
| PR MV\r | |||
| PR VI\n | PR VI\r\n410\r\n> | ||
| PR VM\n | PR VM\r\n6144\r\n> | ||
| VI=410\n | VI=410\r\n> | After Checking for motion / negative limit sensor, response = 'PR MV\r\n1\r\n?' | |
| VM=4096\n | VM=4096\r\n> |
Laser1
| Command | Receive | Parameter | Function |
|---|---|---|---|
| VERSION?\r | SMD-G-1.1.2\r\n\r\n | Read Laser Version. | |
| STAT?\r | ENABLED\r\n | Read laser status. | |
| POWER=p\r | p=[0,430] (milliwatt) | Set laser power. | |
| POWER?\r | pmW\r\n (0000mW\r\n) | p (milliwatt) | Read laser power. |
| ON\r | Turn laser on. | ||
| OFF\r | Turn laser off. | ||
| \n\n |
KLOEHNA
| Command | Parameter | Function |
|---|---|---|
| /1\r | ||
| /1&\r | ||
| /1IR\r | ||
| /1L7R\r | ||
| /1l7R\r | ||
| /1L7R\r | ||
| /1l7R\r | ||
| /1L7R\r | ||
| /1l7R\r | ||
| /1L7R\r | ||
| /1l7R\r | ||
| /1OR\r | ||
| /1V10000A0R\r | ||
| /1W4R\r |
VICIA1
| Command | Parameter | Function |
|---|---|---|
| CC9\r | ||
| CP\r | ||
| CW1\r | ||
| VR\r | ||
| CC1\r | ||
| CW2\r |
[ExcitationFilter1]
FPGACommandSuffix = 1
0 = BlockPosition, Home ; NOTE!!! If spelling is changed, C# code must also be changed!!!
1 = OD4.0
2 = OD2.0
3 = OD0.6
4 = OpenBandPassPosition ; Open. NOTE!!! If spelling is changed, C# code must also be changed!!!
5 = OD0.2
6 = OD1.4
7 = OD1.5
[ExcitationFilter2]
FPGACommandSuffix = 2
0 = BlockPosition,Home ; NOTE!!! If spelling is changed, C# code must also be changed!!!
1 = OD4.5
2 = OD3.0
3 = OD2.0
4 = OpenBandPassPosition ; Open. NOTE!!! If spelling is changed, C# code must also be changed!!!
5 = OD0.2
6 = OD0.9
7 = OD1.0
[ChromaticCompensator]
FPGACommandSuffix = 3
Positions = 0, 399, 799, 1199
0 = Home, Open, Sequencing_AC
1 = Unused1, Sequencing_GT
2 = Unused2
3 = Unused3, Beadchip
- Each filter set represents 4 positions on a wheel
[EmissionFilterSet1] ; This is for SensorPath 1
FPGACommandSuffix = 1
NumPositions = 4
Positions = 0, 399, 799, 1199
0 = Filter.610-60, Home, Blocked
1 = Filter.740-60
2 = Wedge
3 = Open
[EmissionFilterSet2]
FPGACommandSuffix = 2
NumPositions = 4
Positions = 0, 399, 799, 1199
0 = Filter.687-20, Home, Blocked
1 = Filter.558-32
2 = Wedge
3 = Open
VacuumDACSetting= 2500 ; 0 to 4095, HiSeq is 2130
Optical Path
[Analytical Channels]
Channel1 = Grn
Channel2 = Red
[Sequencing Channels]
Channel1 = C
Channel2 = A
Channel3 = T
Channel4 = G
| Path | SensorPath | LaserPath | ExcitationPosition | EmissionPosition | ChromaticCompensatorPosition |
|---|---|---|---|---|---|
| Analytical Channel Grn | 1 | 1 | OpenBandPassPosition | Filter.610-60 | Beadchip |
| Analytical Channel Red | 2 | 2 | OpenBandPassPosition | Filter.687-20 | Beadchip |
| Analytical Channel Reflective | 2 | 2 | OD2.5 Only Ex 2 | Open | Open |
| Analytical Channel Reflective Autocenter | 2 | 2 | OD2.5 Only Ex 2 | Open | Open |
| Analytical Channel Emissive | 1 | 1 | OD1.0 ETF like scanning | Filter.610-60 | Beadchip |
| Sequencing Channel A | 2 | 2 | OpenBandPassPosition | Filter.687-20 | Sequencing_AC |
| Sequencing Channel T | 1 | 1 | OpenBandPassPosition | Filter.610-60 | Sequencing_GT |
| Sequencing Channel G | 2 | 1 | OpenBandPassPosition | Filter.558-32 | Sequencing_GT |
| Sequencing Channel C | 1 | 2 | OpenBandPassPosition | Filter.740-60 | Sequencing_AC |
| Sequencing Channel Reflective | 2 | 2 | OD2.5 Only Ex2 | Open | Open |
| Sequencing Channel Emissive | 1 | 2 | OpenBandPassPosition | Filter.740-60 | Open |
OpenSeq
How about making an Open Source Next Level Sequencing machine - OpenSeq.
Maybe a bit slower and smaller.. like 1 genome per day 🙂
Not sure a price tag of 500k euro is justified for such a machine...
Maybe similar to the iSeq100:
https://www.illumina.com/systems/sequencing-platforms/iseq.html